Is there a function that can map genomic position (hg19) back from a protein position? I can have name of a particular transcript, and exon number.
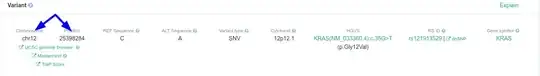
For example, I have KRAS gene for which I would like to know the exact genomic position of G12, and I expect to have an output saying:
chr12, from 25398283 to 25398285.
transvar could be an excellent solution if the current built was working (broken links for fasta annotations). The web search is working but not allowing files heavier than 5kB which is my case.